Brain MRI Segmentation
Utilizing advanced image processing to segment brain tissues
Overview
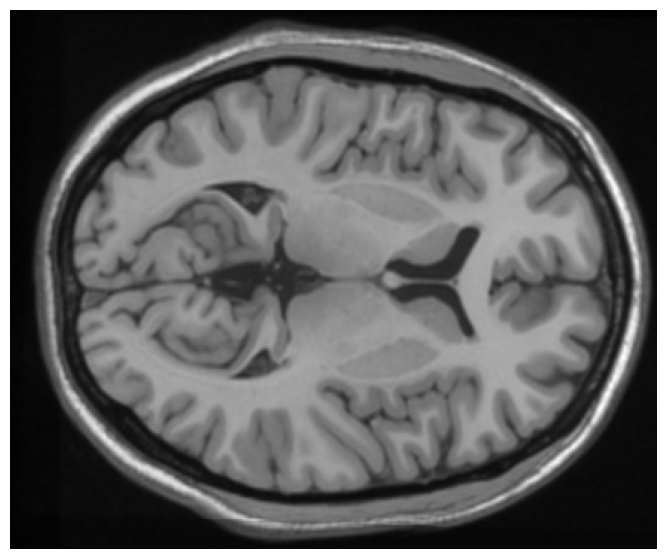
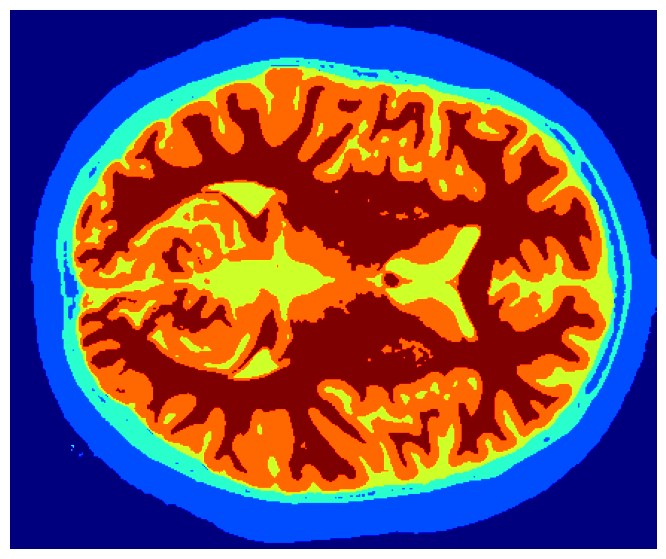
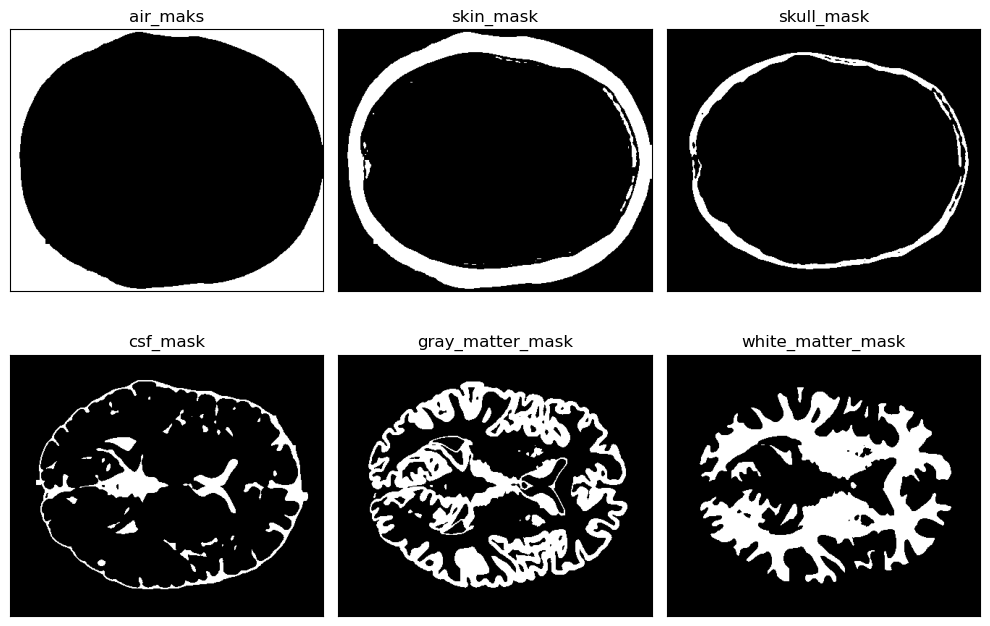
This project tackles the critical task of segmenting brain MRI scans into five distinct tissue layers—background, cerebrospinal fluid (CSF), gray matter, white matter, and skull—using a hybrid approach that merges boundary-based and region-based techniques. The goal was to improve accuracy and computational efficiency for medical applications such as detecting abnormalities, tracking disease progression, and isolating tissues for functional analysis. The pipeline integrates preprocessing (min-max normalization, Gaussian filtering), skull stripping (Otsu’s thresholding, contour detection), and K-Means clustering for tissue classification. Both 2D slices and 3D volumetric data were processed, with the 3D method achieving a 10% speed improvement and superior accuracy (average Dice coefficient: 0.94) by leveraging spatial coherence through connected component labeling. Performance was rigorously validated using overlap metrics like Jaccard Score (IoU) and Dice Coefficient, ensuring robustness across varying noise levels and intensity distributions.
Problem Statement
Accurate segmentation of brain MRI tissues is vital for diagnosing neurological disorders such as Alzheimer’s, tumors, and multiple sclerosis. However, challenges like noise artifacts, intensity inhomogeneity, and overlapping tissue properties (e.g., CSF and gray matter) hinder traditional methods. Existing approaches often suffer from computational inefficiency, poor generalization across slices, or inadequate differentiation of subtle tissue boundaries. This project addresses these limitations by developing a scalable pipeline that optimizes preprocessing, skull stripping, and clustering steps. The solution balances speed and precision, enabling reliable isolation of tissues for clinical workflows while maintaining compatibility with both 2D and 3D MRI datasets.
Methodology
The methodology combines preprocessing, skull stripping, and tissue classification into a cohesive workflow:
- Preprocessing: Raw 2D/3D MRI data undergoes min-max normalization to enhance contrast, followed by Gaussian filtering (σ=1) to reduce noise while preserving edges. The Triangle Threshold algorithm isolates the brain from background air by analyzing intensity histograms. Morphological operations (3x3 kernel for 2D; 3x3x3 kernel for 3D) refine masks by closing gaps and removing noise.
- Skull Stripping: Otsu’s adaptive thresholding generates initial brain and skin masks. Contour detection (via marching squares in 2D) or connected component labeling (in 3D) isolates skin regions, which are subtracted to retain the brain. Morphological closing and
remove_small_holeseliminate residual noise. - Tissue Segmentation: K-Means clustering (k=4) categorizes brain voxels into CSF, gray matter, white matter, and background based on intensity. The 3D method leverages volumetric coherence through connected component labeling, reducing redundant computations across slices.
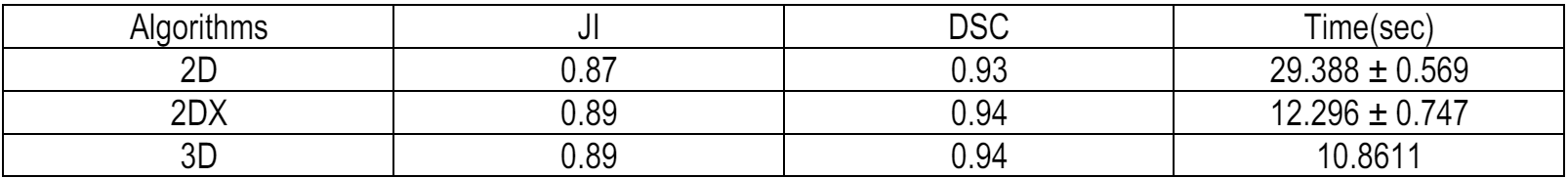
- Evaluation: Performance was assessed using macro-averaged Jaccard Score (IoU) and Dice Coefficient (DSC), ensuring class-agnostic validation. Min-max normalization outperformed alternatives (DSC: 0.93 vs. 0.46 for Z-Score), and kernel size experiments highlighted 3D robustness (DSC: 0.94 with a 3x3x3 kernel).
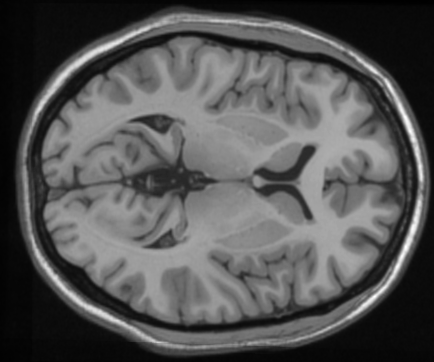
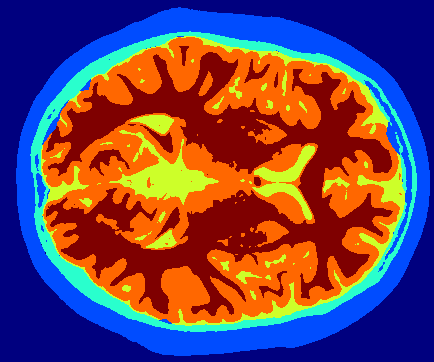
Implementation Details
- Preprocessing:
- Normalization: Min-max scaling was chosen over Z-score due to its superior performance (Table 1: DSC 0.93 vs. 0.46) in enhancing low-contrast tissues.
- Denoising: A Gaussian filter (σ=1) smoothed images without blurring critical boundaries.
- Air Removal: The Triangle Threshold algorithm dynamically identified optimal intensity cutoffs from histograms, outperforming static thresholds.
- Morphological Refinement: Iterative opening/closing operations with 3x3 kernels ensured clean masks.
- Skull Stripping:
- 2D Approach: Marching squares detected skin contours, but slow execution (2.94s/slice) led to the adoption of connected component labeling (1.23s/slice).
- 3D Optimization: Volumetric connected component labeling grouped spatially coherent regions, reducing runtime by 10% (10.86s/volume vs. 2DX’s 12.07s).
- Hole Removal: The
remove_small_holesfunction (area threshold: 100² pixels in 2D; 100²×10 in 3D) addressed residual noise.
- Tissue Classification:
- K-Means Clustering: Intensity-based grouping assigned labels to CSF (lowest intensity), gray matter, white matter, and background.
- 3D Coherence: Volumetric data allowed cross-slice consistency checks, improving accuracy in ambiguous regions.
Evaluation
For evaluation, we utilized the Jaccard Index (JI) and Dice Coefficient (DSC) to measure segmentation accuracy by quantifying the overlap between predicted tissue masks and ground-truth labels. These metrics were macro-averaged across all tissue classes (CSF, gray matter, white matter, skull, and background) to ensure balanced assessment.
Challenges
- Computational Overhead: Initial reliance on 2D contour detection (2.94s/slice) proved impractical for large datasets. Switching to connected component labeling (1.23s/slice) reduced latency by 58%.
- Noise Sensitivity: Residual artifacts in Otsu masks required iterative morphological refinement. The
remove_small_holesfunction and kernel size tuning (3x3 vs. 5x5) were critical for stability. - 3D Parameter Calibration: Adjusting hole thresholds (100²×10) and kernel dimensions (3x3x3) demanded domain expertise to avoid over/under-segmentation.
- Tissue Ambiguity: Intensity overlap between CSF and gray matter occasionally led to misclassification. Future work could integrate texture features or deep learning for finer distinctions.
Technology Stack
- Python
- Scikit-Image
Complete Report
For a more detailed information on the implementation click here
Visualization